MUTATE: IN SILICO PERFORMANCE OF THE FLUCTUATION TEST
Hints
- After download Mutate requires the Java 5.0 (or higher) Runtime Environment freely available at http://www.java.com.
- Whatever the operating system, you should be able to run the program just by double clicking on the mutate.jar file
- WINDOWS USERS: If double-clicking the .jar file does not open the program, make sure that the jar extension is associated with javaw.exe (in your JRE folder).
Contact
For any questions you can contact me
Return to AC-R home
Implementation notes
Mutations occur randomly at each generation under a Poisson distribution with mean Nt * µ * number of defined mutant sites.
In the current implementation the initial number of non-mutant cells N0 is fixed to 1.
Resistance will be acquired when a given mutation, user-defined, is present.
For example if the E. Coli fhuA gene, which encodes phage T1 receptor, is loaded, one or more stop codon mutations can be defined to perform the study.
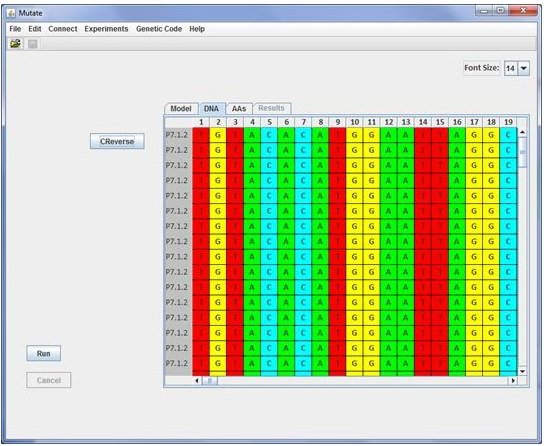
The user should manually define such mutations onto the given DNA sequence before the experiment can be performed. At present two kind of experiments
has been implemented. Experiment 1 (E1) controls that parallel culture and sampling do not produce fluctuation effects in the number of mutations (Luria & Delbruck, 1943).
Experiment 2 (E2) is the fluctuation test. Therefore, in Experiment 1 a single culture is evolved until reaching the user-defined population size (# of bacteria/cc).
Then this culture is divided into equal portions (# of cultures) to be separately tested for phage resistance.
In Experiment 2, a given, user-defined, number of cultures are independently evolved until reach a desired number of bacteria/cc.
Then the cultures are tested for phage resistance.
In each experiment the mean and variance of survival cells per plate are measured.
A. Carvajal-Rodriguez - Departamento de Bioquímica Genética e Inmunología - Universidad de Vigo.
( Last update: December 2011)