SGoFicance Trace web page
Population Genetics and Cytogenetics Group (XB2)
Departamento de Bioquímica, Genética e Inmunología
Área de Genética. Universidad de Vigo
'SGoFicance Trace': assesing significance in high dimensional testing problems
Rationale of the SGoFicance approach
The 'SGoFicance Trace' (from SGoF's significance trace), is a graphical complement to SGoF which can help to make decisions in multiple testing problems. It is constructed from SGoF multitest and uses the same kind of input file. Let SGoF(γ) denote the SGoF algorithm when using a given threshold γ to be tested at α level. The basic idea was to let γ vary on the whole interval (0,1). The SGoFincance Trace displays up to four different plots:
(A) log-significance (p-value) of SGoF(γ) vs. γ
(B) number of effects detected by SGoF(γ) vs. γ
(C) estimated FDR of SGoF(γ) vs. γ
(D) threshold of the pi's vs. γ.
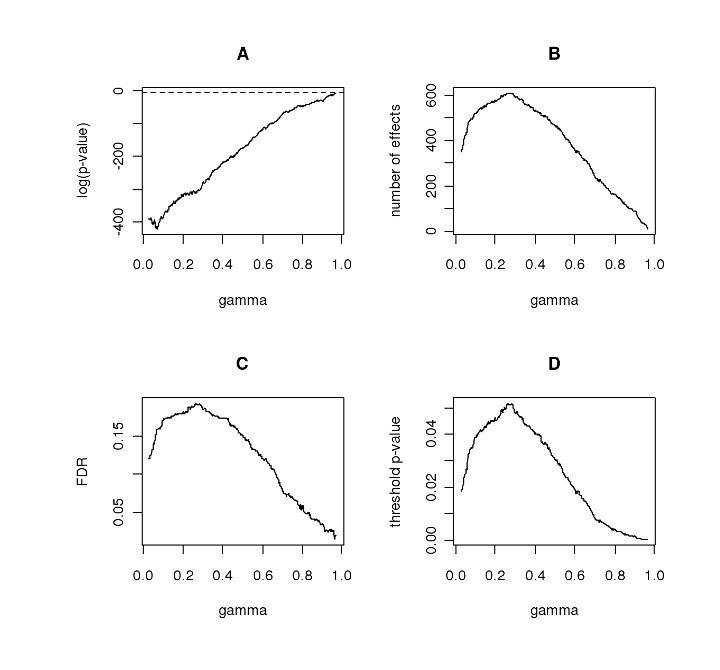
The figure corresponds to the SGoFicance Trace of the data from the microarray study of hereditary breast cancer by Hedenfalk et al. (2001). Also analyzed by Storey et al.(2003). The total number of genes is S = 3,170.
For more detailed explanations please see the papers:
SGoF: BMC Bioinformatics 10:209 (2009) [PDF]
SGoFicance: PLoS ONE 5 (12): e15930 (2010) [PDF]
SGoF+: Carvajal-Rodríguez & de Uña-Alvarez (In preparation)
SGoFicance Trace script
A script in the R language has been written for the computation of the SGoFicance Trace and can be downloaded here .
 2010 Antonio Carvajal Rodríguez. Last updated 18/07/2010
2010 Antonio Carvajal Rodríguez. Last updated 18/07/2010